Embeddings
When you navigate to the Embeddings view of the LightlyOne Platform, you can view the embeddings of all the samples in your dataset in a 2d representation.
On the top of the view, you can choose between three well-known dimensionality reduction methods.
All the samples of your active tag are shown in the scatterplot. This allows you to
pan, by holding your mouse down and moving aroundzoom, by using your mouse wheel
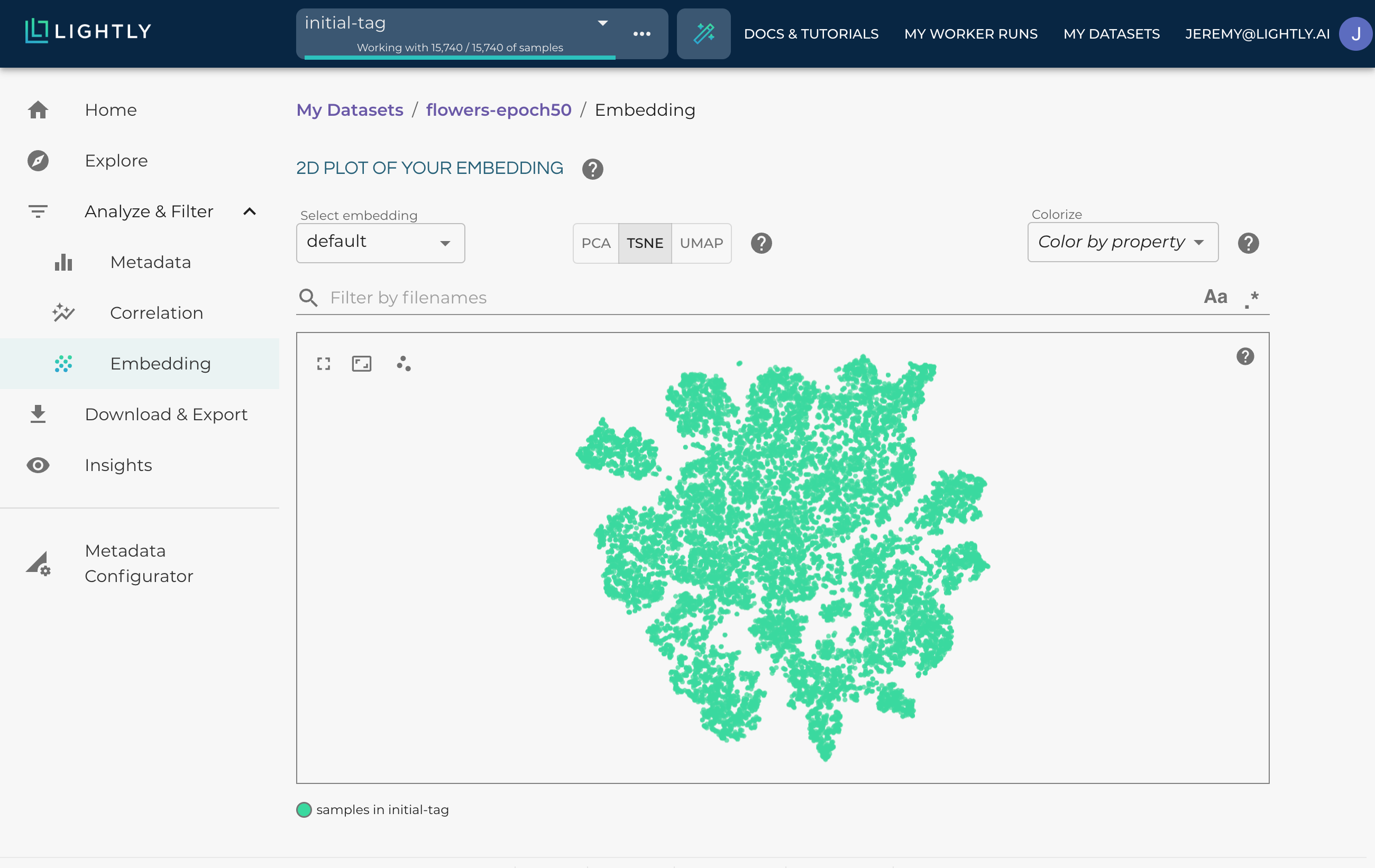
The embedding can show you clusters of your data.
Dimensionality Reduction
LightlyOne uses different dimensionality reduction methods to get a 2d representation of your data.
- PCA: Principal Component Analysis (PCA) is a dimensionality reduction method that is often used to reduce the dimensionality of large datasets so they can be visualized in 2d space. The idea of PCA is simple — reduce the number of variables of a dataset while preserving as much information as possible. For a more thorough explanation, please refer to the PCA article on Wikipedia
- TSNE: t-distributed stochastic neighbor embedding (t-SNE or TSNE) is a dimensionality reduction method that is often used to reduce the dimensionality of large datasets so it can be visualized in 2d space. The idea of TSNE is that nearby points model similar objects, and dissimilar objects are modeled by distant points with high probability. For a more thorough explanation, please refer to the TSNE article on Wikipedia
- UMAP: Uniform Manifold Approximation and Projection (UMAP) is a novel manifold learning technique for dimension reduction. The result is a practical, scalable algorithm that applies to real-world data. The UMAP algorithm is competitive with t-SNE for visualization quality and arguably preserves more of the global structure with superior run-time performance. For a more thorough explanation, please refer to the UMAP article on Wikipedia
Colorize
It's also possible to use the values of a third metadata to either colorize the samples by projecting the range of values on them or categorizing them to show/hide them as legends.
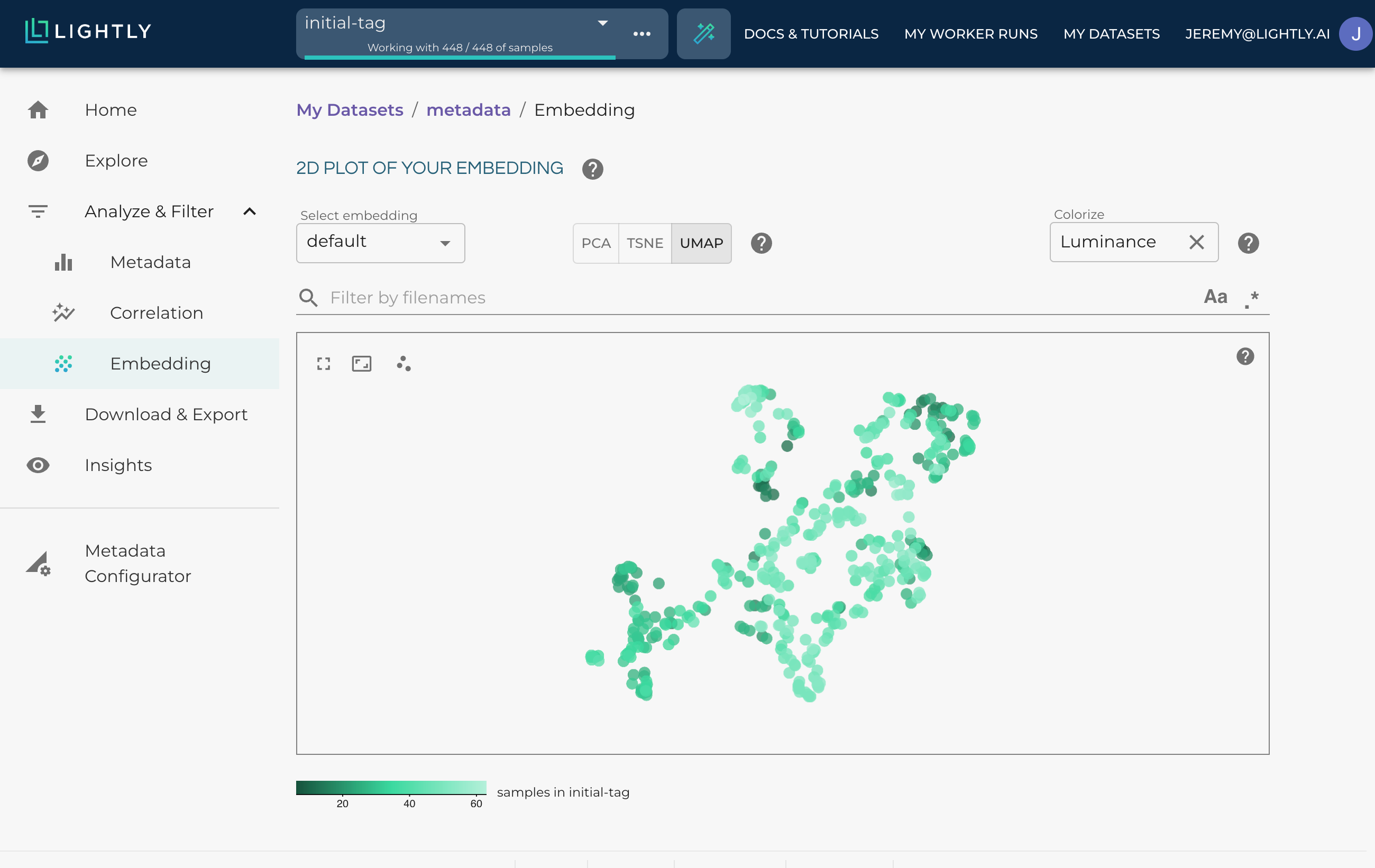
Using a numerical metadata to project the range of values.
Select Clusters
Once interesting clusters or outliers are detected, selecting them for further inspection is possible.
You can select a cluster by pressing the shift key and holding the mouse down simultaneously. This will activate the lasso modus, with which you can select the samples of interest by drawing a circle.
If you have multiple clusters you want to inspect simultaneously, you can press the shift and alt keys while holding the mouse down to select multiple regions.
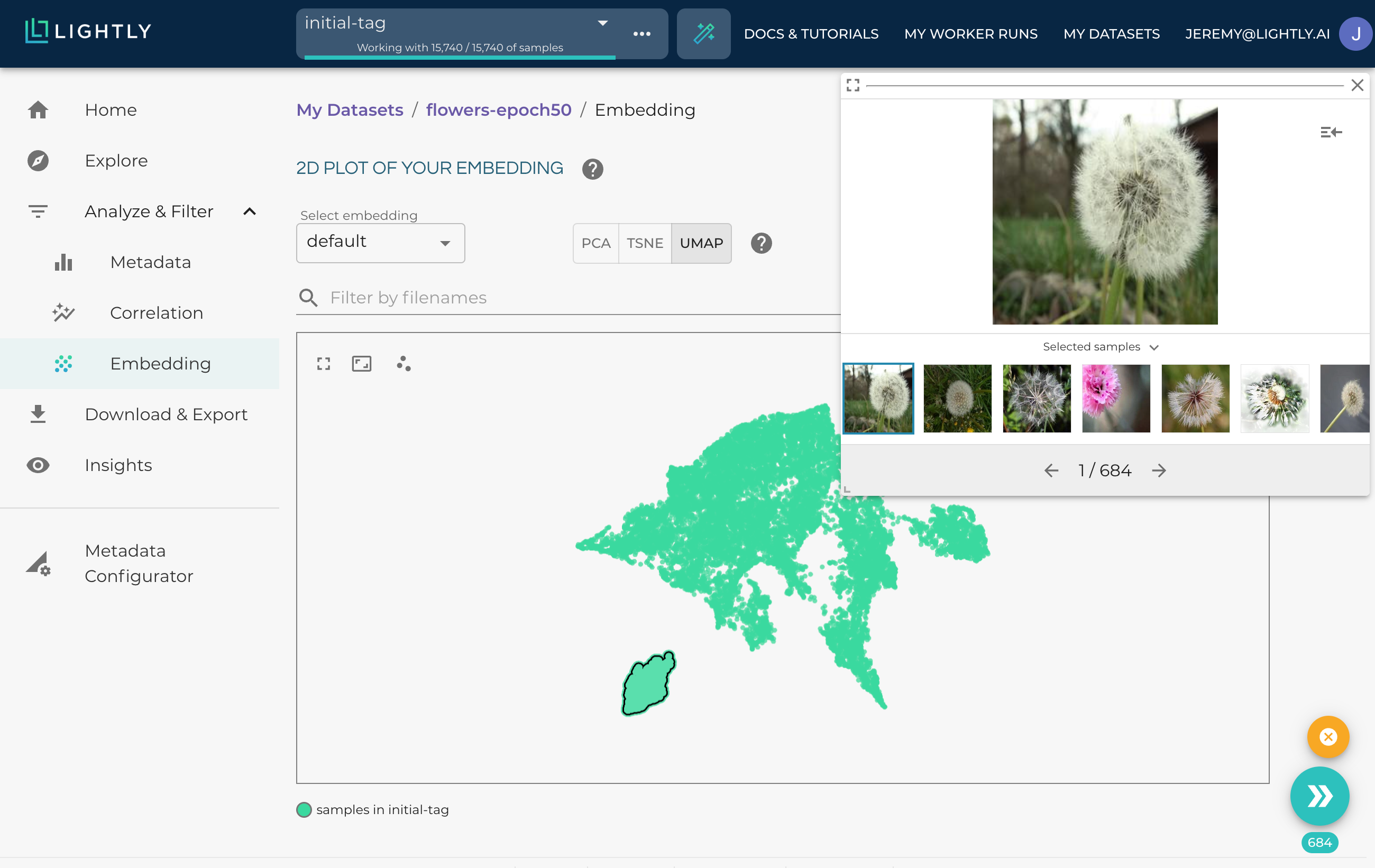
Inspecting a cluster is very easy by pressing the shift-key and drawing a circle.
Updated 10 months ago
