Work with Metadata
LightlyOne can make use of metadata collected alongside your images or videos. Provided metadata can be used to steer the selection process and to analyze the selected dataset in the LightlyOne Platform
Metadata Folder Structure
Following, we outline the format in which metadata can be added to a Lightly datasource. Everything regarding metadata takes place in a subdirectory of your configured Lightly datasource called .lightly/metadata. Below we show an example structure of an input datasource containing images and a Lightly datasource containing the metadata files:
input_datasource/
├── image_0.png
└── subdir/
├── image_1.png
├── image_2.png
├── ...
└── image_N.png
lightly_datasource/
└── .lightly/metadata/
├── schema.json
├── image_0.json
└── sudir/
├── image_1.json
├── image_2.json
├── ...
└── image_N.jsonIt is important that the relative filename path of the images to the input_datasource is exactly the same as the relative filename path of the predictions to the .lightly/metadata directory. This way, LightlyOne can match the predictions to the correct source images or videos. As an example, the
prediction file for input_datasource / subdir/image_1.png should be at lightly_datasource/.lightly/metadata / subdir/image_1.json. Note that the subdir part of the path is
included in both cases.
All of the .json files are explained in the following sections.
Changing the Input datasourceMake sure to also use a new Lightly datasource whenever you use a new Input datasource. Otherwise the relative filename path changes and thus the metadata are not found anymore. For details, see Datasource Configuration for Multiple Projects. If you want to use one or multiple subdirectories, e.g. the
subdir, as input, then keep theinput_datasourceunchanged and use the relevant filenames feature.
Check out our reference projectIf you want to have a look at a reference project with the proper folder structure to work with predictions and metadata we suggest having a look at our example here: https://github.com/lightly-ai/object_detection_example_structure
Metadata Schema
The schema defines the metadata format and helps the LightlyOne Platform and Worker correctly identify and display different types of metadata. You can provide this information to LightlyOne by adding a schema.json file to the .lightly/metadata directory in your Lightly datasource. The schema.json file must contain a list of configuration entries. Each of the entries is a dictionary with the following keys:
name: Name of this metadata item (can be chosen by the user)path: Concatenation of the keys of the metadata item, for example,myObject.myKey1.myKey2. We reserve the top-level keylightlyfor internal use, please make sure your metadata does not use it at the top level.defaultValue: Fallback value if a sample has no corresponding metadata entryvalueDataType: Data type, one of the following:CATEGORICAL_STRINGCATEGORICAL_BOOLEANCATEGORICAL_INTNUMERIC_FLOATNUMERIC_INT
For example, let’s say we have additional information about the scene and weather for each image we have collected:
{
"scene": "Highway",
"weather": {
"description": "sunny",
"temperature": 27.2,
"air_pressure": 1025
},
"vehicle_id": 0
}A possible schema could look like this:
[
{
"name": "Scene",
"path": "scene",
"defaultValue": "undefined",
"valueDataType": "CATEGORICAL_STRING"
},
{
"name": "Weather description",
"path": "weather.description",
"defaultValue": "nothing",
"valueDataType": "CATEGORICAL_STRING"
},
{
"name": "Temperature",
"path": "weather.temperature",
"defaultValue": 0.0,
"valueDataType": "NUMERIC_FLOAT"
},
{
"name": "Air pressure",
"path": "weather.air_pressure",
"defaultValue": 0,
"valueDataType": "NUMERIC_INT"
},
{
"name": "Vehicle ID",
"path": "vehicle_id",
"defaultValue": 0,
"valueDataType": "CATEGORICAL_INT"
}
]Metadata Files
LightlyOne requires a single metadata file per image, video, or frame. LightlyOne assumes the default value from the schema.json if no metadata file exists for an input file. If a metadata file is provided for a full video, LightlyOne assumes that the metadata is valid for all frames in that video.
To provide metadata for an image or a video, place a metadata file with the same name as the image or video in the .lightly/metadata directory in the Lightly bucket but change the file extension to .json. The file should contain the metadata in the format defined under Metadata Format (we will get there).
# Filename of the metadata for file in input_datasource/FILENAME.EXT
lightly_datasource/.lightly/metadata/${FILENAME}.json
# Example for image in input_datasource/subdir/image_1.png
lightly_datasource/.lightly/metadata/subdir/image_1.json
# Example for image in input_datasource/image_0.png
lightly_datasource/.lightly/metadata/image_0.json
# Example for video in input_datasource/subdir/video_1.mp4
lightly_datasource/.lightly/metadata/subdir/video_1.jsonWhen working with videos, providing a metadata file per frame is possible. LightlyOne assumes the default value from the schema.json if a frame has no corresponding metadata file. The metadata file name must contain the original video name, video extension, and frame number in the following format:
{VIDEO_NAME}-{FRAME_NUMBER}-{VIDEO_EXTENSION}.json
Frame numbers start from 0. They are zero-padded to the total length of the number of frames in a video. A video with 200 frames must have the frame number padded to length three. For example, the frame number for frame 99 becomes 099. A video with 1000 frames must have frame numbers padded to length four (99 becomes 0099). You can use this python code snippet to set the filename:
zero_padding = len(str(len(video_frames)))
metadata_filenames[frame_index] = f"{video_name}-{frame_index:0{zero_padding}}-{video_extension}.json"Examples are shown below:
# Filename of the metadata of the Xth frame of video input_datasource/FILENAME.EXT
# with 200 frames (padding: len(str(200)) = 3)
lightly_datasource/.lightly/metadata/${FILENAME}-${X:03d}-${EXT}.json
# Example
# Video: input_datasource/subdir/video_1.mp4
# Metadata file for frame 99 of 200 frames:
lightly_datasource/.lightly/metadata/subdir/video_1-099-mp4.json
# Example
# Video: input_datasource/video_0.mp4
# Metadata file for frame 99 of 200 frames:
lightly_datasource/.lightly/metadata/subdir/video_0-099-mp4.jsonMetadata Format
Metadata for images and videos require the keys type, and metadata. Here, type indicates whether the metadata is per image, video, or frame. And metadata contains the actual metadata.
Image
{
"type": "image",
"metadata": {
"scene": "highway",
"weather": {
"description": "rainy",
"temperature": 10.5,
"air_pressure": 1
},
"vehicle_id": 321
}
}Video
{
"type": "video",
"metadata": {
"scene": "city street",
"weather": {
"description": "sunny",
"temperature": 23.2,
"air_pressure": 1
},
"vehicle_id": 321
}
}Frame
{
"type": "frame",
"metadata": {
"scene": "city street",
"weather": {
"description": "sunny",
"temperature": 23.2,
"air_pressure": 1
},
"vehicle_id": 321
}
}Crop
Custom metadata for object crops is currently not supported.
Next Steps
If metadata is provided, the LightlyOne Worker will automatically detect and load it into the LightlyOne Platform, where it can be visualized and analyzed after running a selection.
Visualizing the different metadata categories in the LightlyOne Platform embedding view is also possible. The following example shows the categorical metadata “Scene” from the BDD100k dataset:
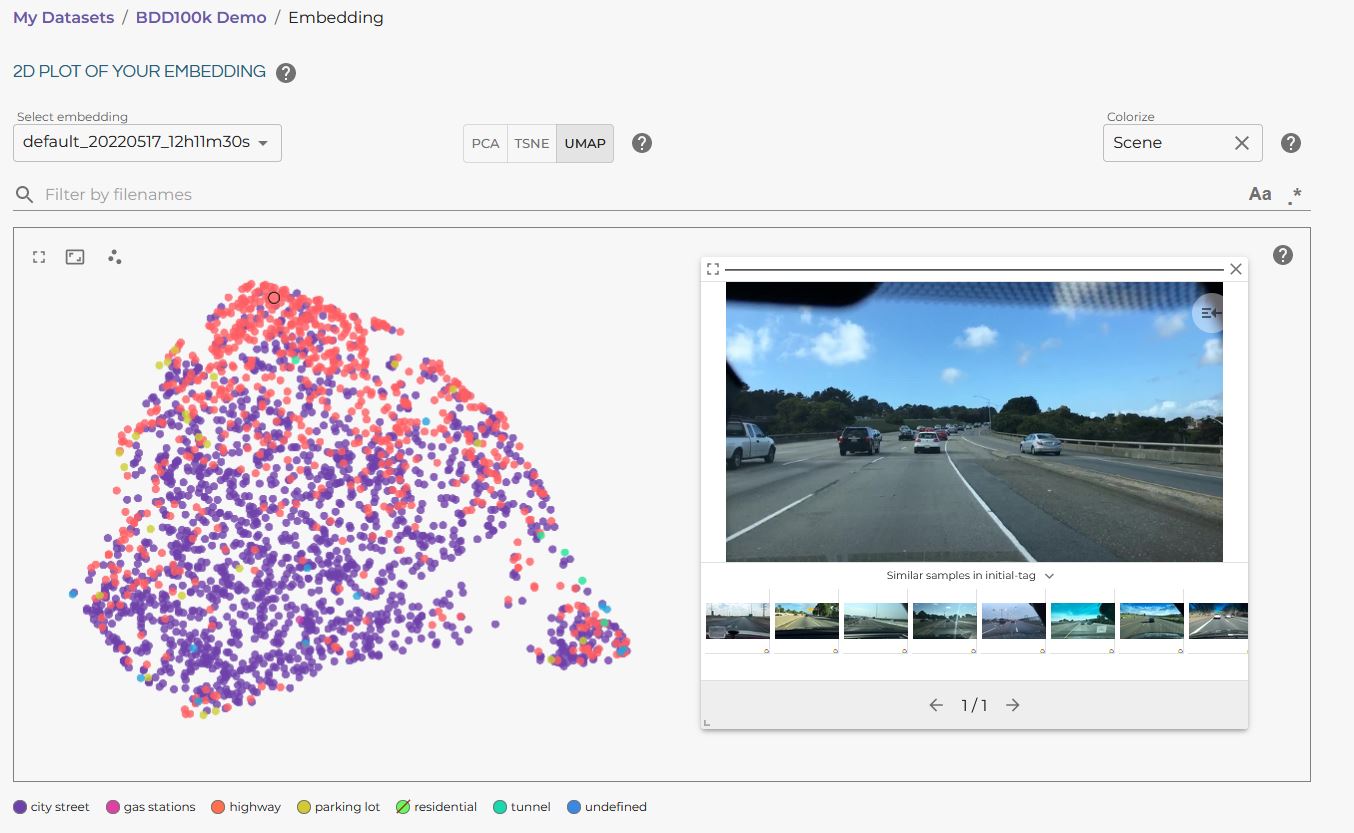
BDD100K embedding visualization in the LightlyOne Platform.
For information on how to use metadata for selection, head to Customize a Selection and the section on Metadata input.
Updated 6 months ago
